
Resistance to β-lactam antibiotics mediated by alterations of PBPs has been reported in some gram-negative bacteria.

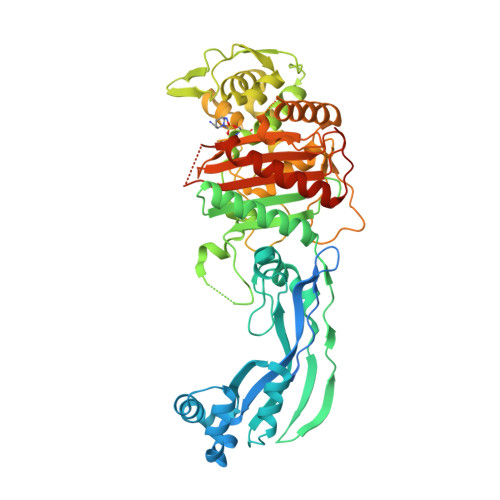
These enzymes are inserted in the cytoplasmic membrane only at their amino termini, and water-soluble forms have been obtained that should be suitable for crystallization and X-ray analysis. The high-molecular-weight PBPs of Escherichia coli are believed to possess an amino-terminal peptidoglycan transglycosylase domain and a carboxy-terminal penicillin-sensitive transpeptidase domain. This view is strongly supported by the recent finding of a similarity in the three-dimensional structures of a low-molecular-weight PBP and class A β-lactamases. The availability of the amino acid sequences of several low-molecular-weight PBPs, high-molecular-weight PBPs, and active-site serine β-lactamases has provided evidence that these groups of enzymes have a common, but distant, evolutionary origin. and class B PBPs (PBP1a/b, PBP2, and PBP3) is typically lethal. It might play an important role in the positioning of these proteins within the divisome.Β-Lactam antibiotics exert their antibacterial effects by inactivating the high-molecular-weight penicillin-binding proteins (PBPs) that are responsible for the final stages of peptidoglycan biosynthesis. spectrum activity against class A, C and D -lactamases.
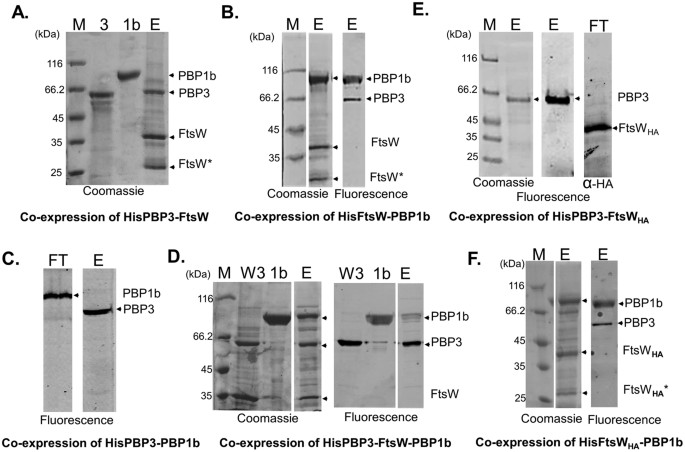
The periplasmic loop 9/10 of FtsW appeared to be involved in the interaction with both PBP1B and PBP3. amino acid changes near the active site of PBP3, the primary target of sulbactam. The HMW-PBPs group can be subdivided into classes A and B, the LMW group into classes A, B and C. PBP1a and PBP1b are involved in both cell elongation and division ( Bertsche et al., 2006 ), whereas the role of PBP1c is likely to be involved in a PG repair mechanism ( Budd et al., 2004 ). However, it could not be detected in the immunoprecipitated FtsW-PBP3 complex. coli, the C-terminal domain of both class A PBPs (PBP1a, PBP1b, and PBP1c) and the class B PBPs (PBP2 and PBP3) (Sauvage et al., 2008) have TPase activity. By using a two-hybrid assay, the class A PBP1B was shown to interact with FtsW. For example Staphylococcus aureus PBP2 is a class A PBP similar to E. The K2-V42 peptide of PBP3 containing the membrane-spanning sequence is a structural determinant sufficient for interaction with FtsW and for PBP3 dimerization. These proteins are able to form a discrete complex independently of the other cell-division proteins. In this work we show direct interactions between FtsW and PBP3 in vivo and in vitro by FRET (Förster resonance energy transfer) and co-immunoprecipitation experiments. CAZ-AVI susceptible KPC-Kp strains became resistant when complemented with mutated bla KPC genes. It requires the putative lipid II flippase FtsW for its localization at the division site and is necessary for the midcell localization of the class A PBP1B. analysis for the original and derived mutants identied CAZ-AVI resistance-associated mutations in KPCs, PBP3 (encoded by ftsI), and LamB, an outer membrane maltoporin. In Escherichia coli, the class B PBP3 is specific for septal peptidoglycan synthesis. The morphogenetic protein complex responsible for the septation during cell division (the divisome) includes class A and class B penicillin-binding proteins (PBPs). During the cell cycle of rod-shaped bacteria, two morphogenetic processes can be discriminated: length growth of the cylindrical part of the cell and cell division by formation of two new cell poles.
